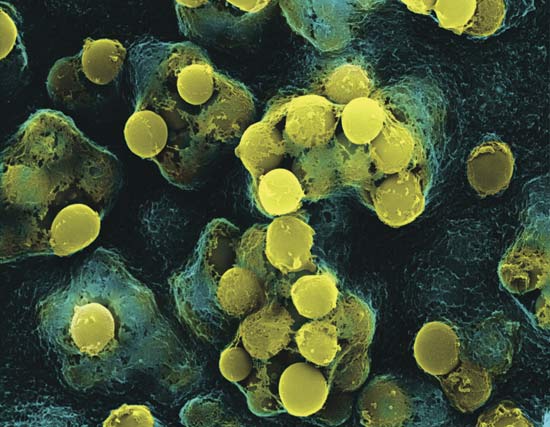
The genomes of hundreds of bacterial strains that cause pneumonia have been sequenced and may lead to new antibiotics and vaccines.
240 lineages of multidrug resistant Streptococcus pneumoniae were collected from around the world and their genomes sequenced in order to understand how the bacteria came to be so virulent.
The research, published in the journal Science this week, compared the genetic sequences with the geographic locations of each specimen to produce a map of the major evolutionary events that have led to the diversity we see today. The team of scientists also pinpointed Europe as the probable birthplace of the first multidrug resistant individual.
The researched also identified recombination as the dominant mechanism the bacteria have used to evolve resistance to antibiotic drugs. Recombination involves individual pieces of DNA moving around the bacterial genome, and in doing so creates new genes. Some of these recombination will result in the bacteria becoming resistant. Many of these drug resistant genes can pass horizontally, from bacteria to bacteria, and could explain how drug resistance has spread so quickly across the globe.
4 million cases of fatal pneumococcal disease are reported each year, and, according to the World Health Organisation, is responsible for an estimated 18% of all deaths of children under the age of 5.
The research was created thanks to a partnership between the Sanger Institute, a world leader in genomic analysis, and scientists from Rockafeller University studying the patterns of illness around the world. Alexander Tomaz, co-author of the paper, praised the unusual collaboration. “Such an alliance between molecular biology and epidemiology promises further interesting insights into the mechanism of bacterial evolution”.
Professor Brian Spratt, a molecular epidemiologist at Imperial highlighted the importance of this research, saying: “how bacteria diversify over the very short time scales […] are of crucial importance for understanding and predicting the response of pathogens to new antibiotics and vaccines.”